# Participant information
participants <- tibble(
participant_id = 1:4,
age = c(22, 25, 19, 31),
condition = c("Control", "Treatment", "Treatment", "Control")
)
# Survey responses (collected separately)
survey <- tibble(
participant_id = c(1, 2, 3, 5), # Note: 4 is missing, 5 is extra
depression = c(12, 18, 10, 25),
anxiety = c(15, 20, 12, 30)
)Joins
PSY 410: Data Science for Psychology
2026-05-11
Why join data?
Three files, one question
Your participant took a survey on Monday and did a cognitive task on Tuesday. The survey data is in one CSV. The task data is in another. Their demographics are in a third.
You want to ask: “Did anxious participants respond more slowly?”
You can’t answer that from any single file. You need joins — the tool that connects separate tables through a shared key.
Two tables, one question
The two tables
Participants:
How do we combine these based on participant_id?
Keys
What are keys?
Keys are variables that uniquely identify observations and connect tables.
Primary key: Uniquely identifies each observation in a table
participant_idin the participants table- Each participant appears only once
Foreign key: References the primary key of another table
participant_idin the survey table- Links back to the participants table
Checking for unique keys
Before joining, verify that your key truly identifies each row:
# A tibble: 0 × 2
# ℹ 2 variables: participant_id <int>, n <int>Empty result = good! Each participant appears only once.
When keys aren’t unique
Sometimes keys aren’t unique (e.g., longitudinal data):
Mutating joins
The join family
Mutating joins add columns from one table to another:
| Function | Keeps rows from… |
|---|---|
left_join() |
Left table (all rows) |
right_join() |
Right table (all rows) |
inner_join() |
Both tables (only matches) |
full_join() |
Both tables (all rows) |
left_join(): Most common
Keep all rows from the left table, add matching data from the right:
# A tibble: 4 × 5
participant_id age condition depression anxiety
<dbl> <dbl> <chr> <dbl> <dbl>
1 1 22 Control 12 15
2 2 25 Treatment 18 20
3 3 19 Treatment 10 12
4 4 31 Control NA NA- Participant 4 has no survey data →
NA - Participant 5 from survey isn’t in participants → excluded
Visualizing left_join()
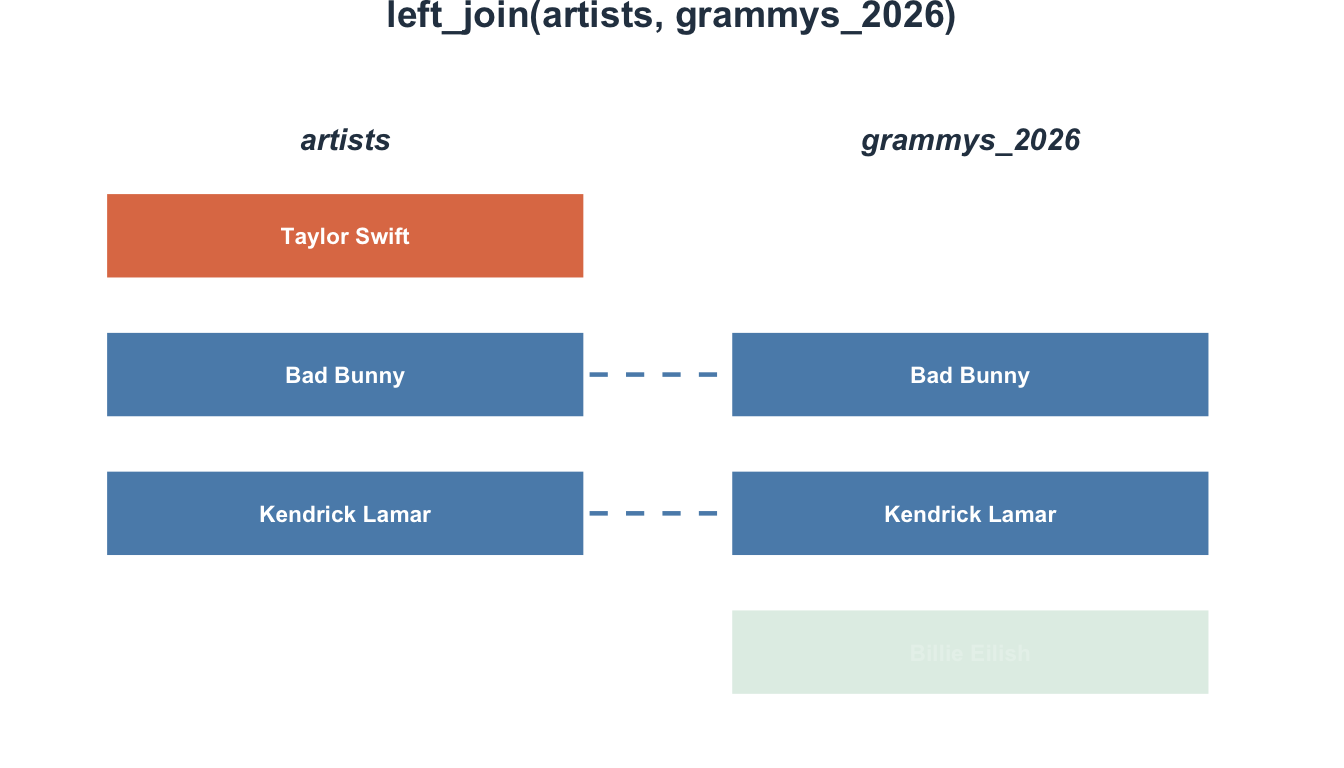

right_join(): The mirror
Keep all rows from the right table:
# A tibble: 4 × 5
participant_id age condition depression anxiety
<dbl> <dbl> <chr> <dbl> <dbl>
1 1 22 Control 12 15
2 2 25 Treatment 18 20
3 3 19 Treatment 10 12
4 5 NA <NA> 25 30Now participant 5 is included (with NA for age/condition), but participant 4 is excluded.
Visualizing right_join()
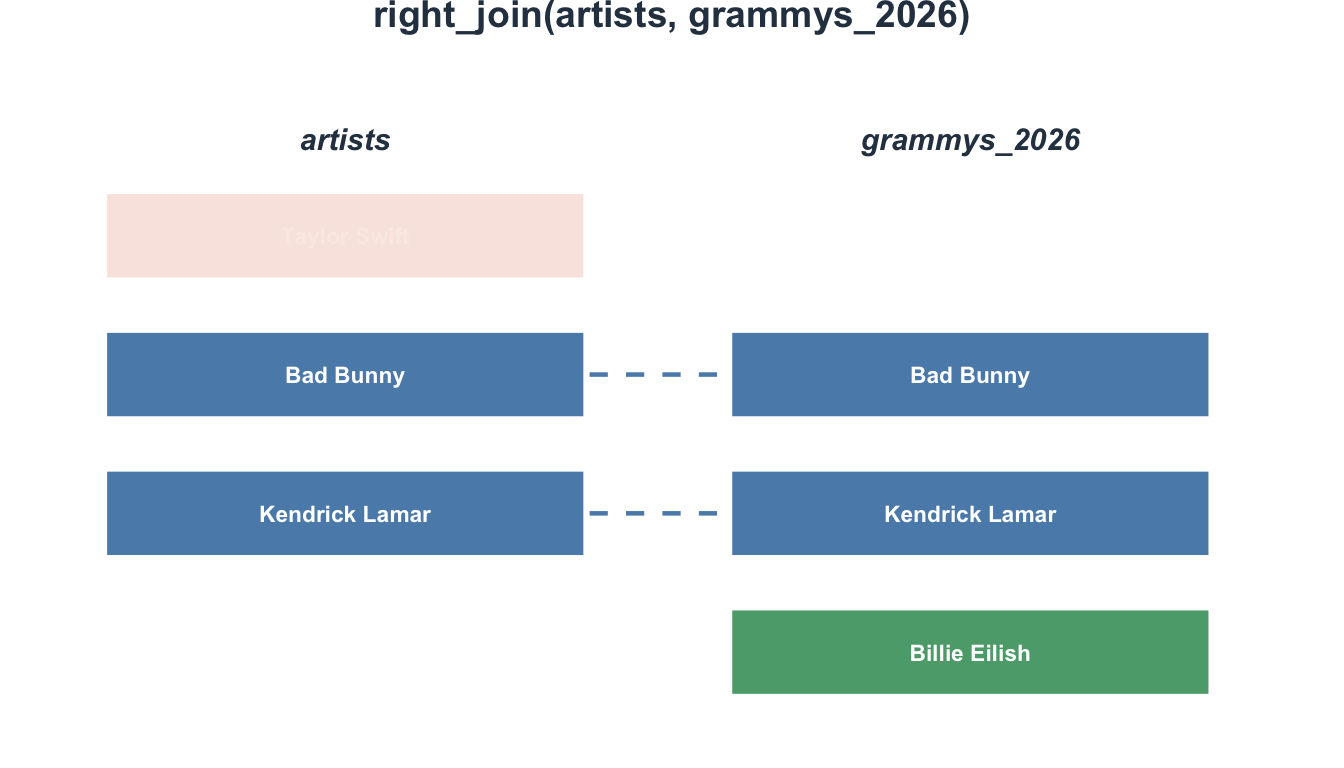

inner_join(): Only matches
Keep only rows that exist in both tables:
# A tibble: 3 × 5
participant_id age condition depression anxiety
<dbl> <dbl> <chr> <dbl> <dbl>
1 1 22 Control 12 15
2 2 25 Treatment 18 20
3 3 19 Treatment 10 12- Participant 4 excluded (no survey data)
- Participant 5 excluded (not in participants)
Visualizing inner_join()
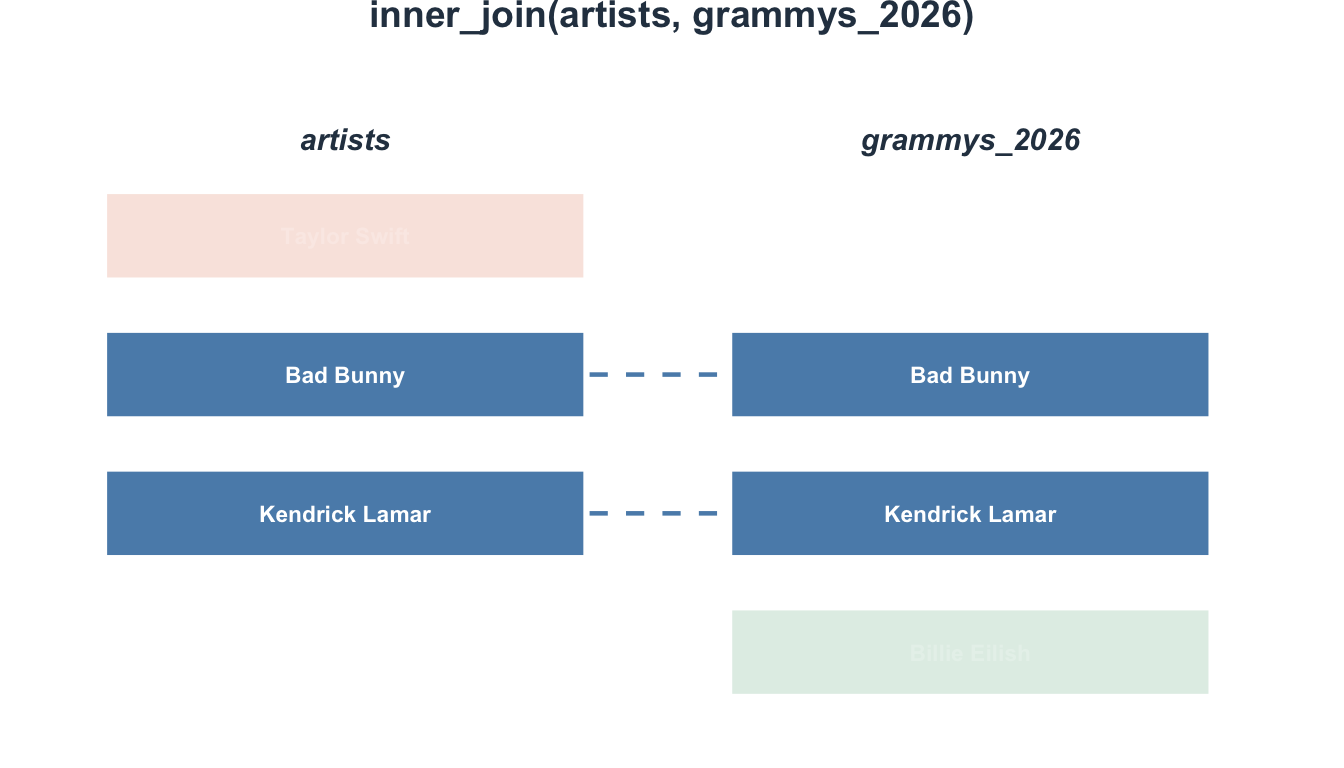

full_join(): Everything
Keep all rows from both tables:
# A tibble: 5 × 5
participant_id age condition depression anxiety
<dbl> <dbl> <chr> <dbl> <dbl>
1 1 22 Control 12 15
2 2 25 Treatment 18 20
3 3 19 Treatment 10 12
4 4 31 Control NA NA
5 5 NA <NA> 25 30Every participant appears, with NA where data is missing.
Visualizing full_join()
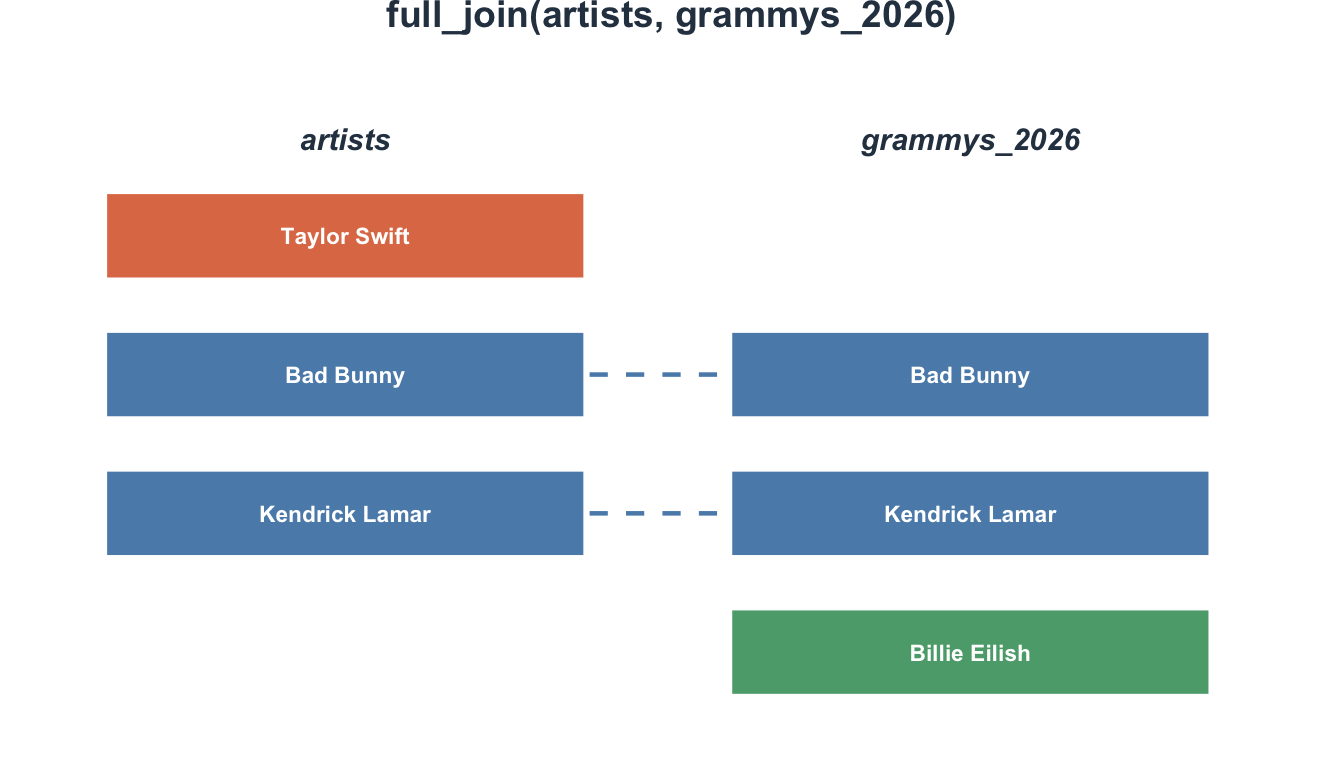

Join types at a glance
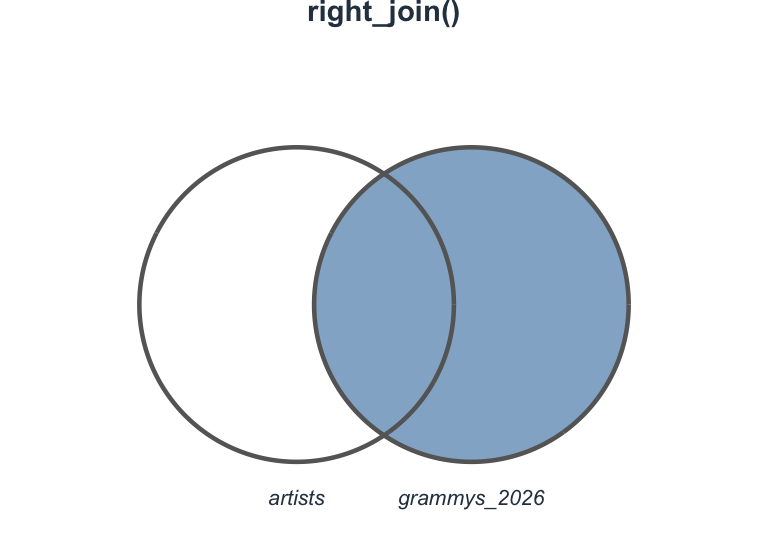
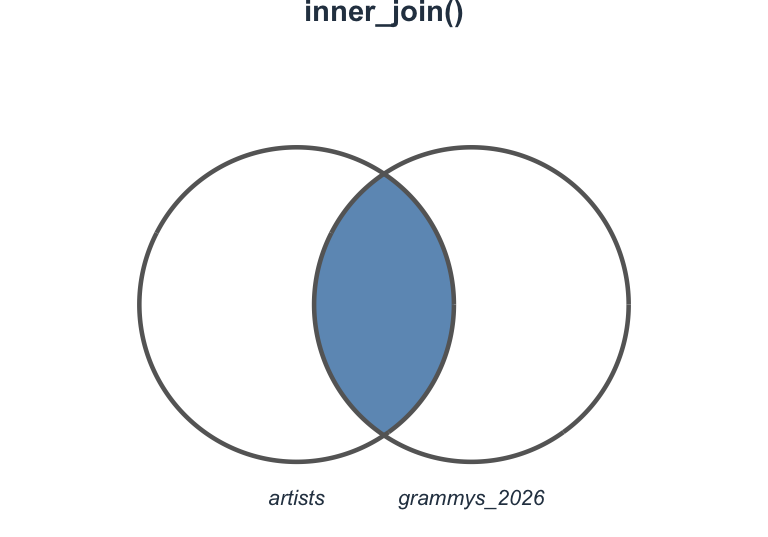
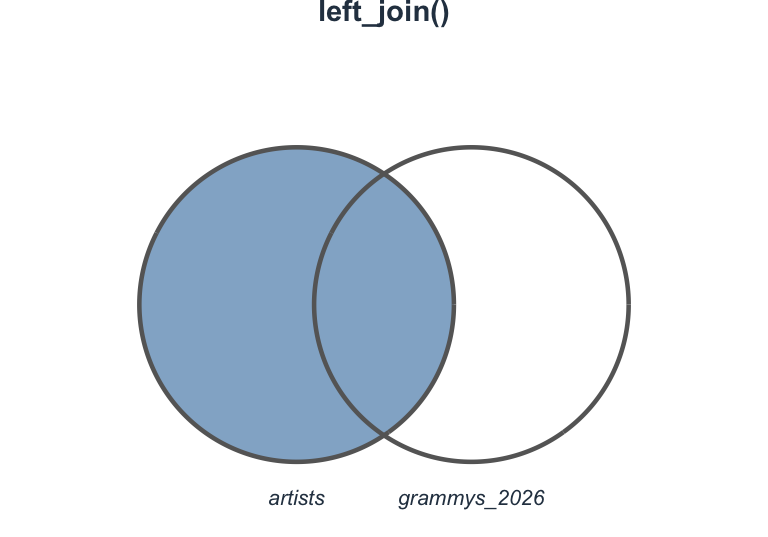

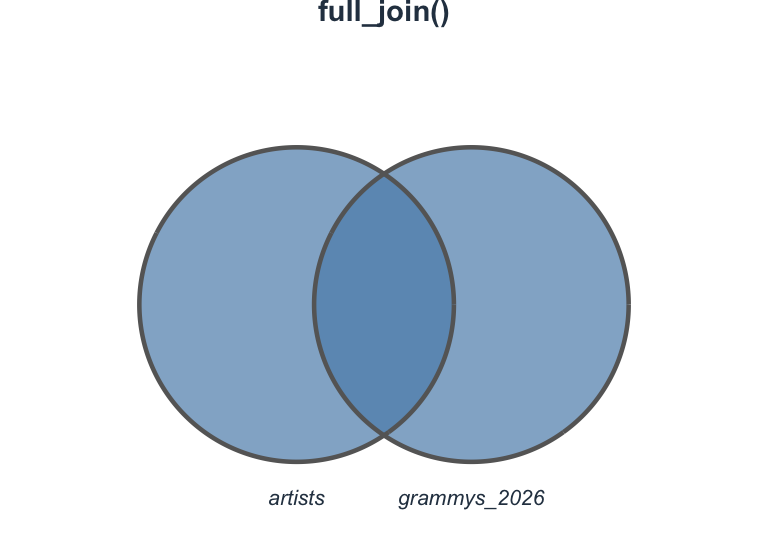
Which join should I use?
Most common in psychology: left_join()
Use when:
- You have a main participant table
- You want to add additional data (surveys, behavioral tasks)
- You want to keep all participants, even those with missing data
Tip
Start with left_join() unless you have a specific reason to use another type.
Psychology example: Adding demographics
# Task data
task_data <- tibble(
participant_id = c(1, 2, 3, 1, 2, 3),
trial = c(1, 1, 1, 2, 2, 2),
reaction_time = c(450, 380, 520, 430, 395, 510)
)
# Add participant info to each trial
task_data |>
left_join(participants, by = "participant_id")# A tibble: 6 × 5
participant_id trial reaction_time age condition
<dbl> <dbl> <dbl> <dbl> <chr>
1 1 1 450 22 Control
2 2 1 380 25 Treatment
3 3 1 520 19 Treatment
4 1 2 430 22 Control
5 2 2 395 25 Treatment
6 3 2 510 19 TreatmentJoining by multiple keys
For longitudinal data, join on multiple columns:
Combining longitudinal data
# Combine into one table
all_times <- bind_rows(time1, time2)
# Add baseline demographics
all_times |>
left_join(
participants |> select(participant_id, age, condition),
by = c("id" = "participant_id") # Keys have different names
)# A tibble: 6 × 5
id time depression age condition
<dbl> <dbl> <dbl> <dbl> <chr>
1 1 1 20 22 Control
2 2 1 18 25 Treatment
3 3 1 25 19 Treatment
4 1 2 15 22 Control
5 2 2 12 25 Treatment
6 3 2 22 19 TreatmentPair coding break
Your turn: Combine study data
You have three tables from a therapy study:
baseline <- tibble(
id = 1:5,
age = c(25, 30, 22, 35, 28),
baseline_depression = c(22, 25, 18, 30, 20)
)
treatment <- tibble(
id = c(1, 2, 3, 4), # Missing id 5
condition = c("CBT", "Control", "CBT", "Control")
)
followup <- tibble(
id = c(1, 2, 3, 5), # Missing id 4, but has id 5
followup_depression = c(12, 23, 10, 18)
)Your turn: Combine study data
- Create a dataset with baseline info and treatment condition (keep all participants)
- Add followup data (keep all from step 1)
- How many participants are missing followup data?
Time: 10 minutes
Filtering joins
A different purpose
Filtering joins don’t add columns — they filter rows based on whether matches exist:
semi_join()— Keep rows that have a matchanti_join()— Keep rows that don’t have a match
semi_join(): Has a match
Keep rows from the left table that exist in the right table:
# A tibble: 3 × 3
participant_id age condition
<int> <dbl> <chr>
1 1 22 Control
2 2 25 Treatment
3 3 19 TreatmentOnly participants 1, 2, and 3 completed the survey.
anti_join(): No match
Keep rows from the left table that don’t exist in the right table:
# Which participants didn't complete the survey?
participants |>
anti_join(survey, by = "participant_id")# A tibble: 1 × 3
participant_id age condition
<int> <dbl> <chr>
1 4 31 Control Participant 4 didn’t complete the survey.
Psychology use case: Attrition analysis
# Who was enrolled
enrolled <- tibble(
participant_id = 1:10,
condition = rep(c("Treatment", "Control"), each = 5))
# Who completed
completed <- tibble(
participant_id = c(1, 2, 3, 4, 7, 8, 9, 10)) # Missing 5 and 6
# Who dropped out?
enrolled |>
anti_join(completed, by = "participant_id")# A tibble: 2 × 2
participant_id condition
<int> <chr>
1 5 Treatment
2 6 Control Attrition by condition
# A tibble: 2 × 2
condition n
<chr> <int>
1 Control 1
2 Treatment 1Warning
Both dropouts were from the Treatment condition — this could bias results!
Common join problems
Problem 1: Keys with different names
Problem 2: Duplicate keys
What happens with duplicates?
# A tibble: 4 × 3
id age score
<dbl> <dbl> <dbl>
1 1 25 85
2 1 25 85
3 2 30 90
4 3 22 88Each duplicate gets matched → creates extra rows!
Important
Always check for duplicate keys before joining.
Problem 3: Missing keys
Joining with NAs
# A tibble: 4 × 3
id response age
<dbl> <chr> <dbl>
1 1 Yes 25
2 2 No 30
3 NA Yes NA
4 3 No 22Row with NA id can’t match → gets NA for age.
Fix the data before joining!
Problem 4: Many-to-many relationships
Many-to-many result
# A tibble: 8 × 5
id trial rt session mood
<dbl> <dbl> <dbl> <dbl> <dbl>
1 1 1 450 1 5
2 1 1 450 2 6
3 1 2 430 1 5
4 1 2 430 2 6
5 2 1 380 1 4
6 2 1 380 2 5
7 2 2 390 1 4
8 2 2 390 2 5Creates all combinations — probably not what you want!
Solution: Be more specific about what you’re joining on, or reshape first.
Practical psychology examples
Example 1: Qualtrics + demographics
# Survey responses exported from Qualtrics
qualtrics <- tibble(
ResponseId = c("R_1", "R_2", "R_3"),
PHQ9_total = c(12, 18, 8),
GAD7_total = c(10, 15, 6)
)
# Demographics collected separately
demo <- tibble(
ResponseId = c("R_1", "R_2", "R_3"),
age = c(25, 30, 22),
gender = c("Female", "Male", "Female")
)Example 1: Result
Example 2: Pre-post intervention
Example 2: Result
Example 3: Item-level to scale-level
Example 3: Result
End-of-deck exercise
Practice: Longitudinal study
You have data from a three-wave longitudinal study:
# Baseline demographics
baseline_demo <- tibble(
pid = 1:6,
age = c(20, 22, 25, 19, 24, 21),
condition = rep(c("Intervention", "Control"), each = 3)
)
# Time 1 assessment
time1_scores <- tibble(
pid = 1:6,
depression_t1 = c(22, 25, 18, 20, 24, 19)
)
# Time 2 assessment (2 dropouts)
time2_scores <- tibble(
pid = c(1, 2, 3, 4, 6), # Missing 5
depression_t2 = c(15, 24, 12, 18, 17)
)
# Time 3 assessment (1 more dropout)
time3_scores <- tibble(
pid = c(1, 2, 3, 6), # Missing 4 and 5
depression_t3 = c(12, 22, 10, 15)
)Your tasks
- Create a complete dataset with all timepoints
- Compute change scores (T1 to T3)
- Identify who dropped out at each wave
- Check if dropout differs by condition
Wrapping up
Join decision flowchart
Ask yourself:
- Do I want to add columns?
- Yes → Use a mutating join
- No → Use a filtering join
- Which rows do I want to keep?
- All from left →
left_join() - All from right →
right_join() - Only matches →
inner_join() - Everything →
full_join()
- All from left →
- Am I filtering rows?
- Keep matches →
semi_join() - Keep non-matches →
anti_join()
- Keep matches →
Key takeaways
- Joins combine data from multiple tables using key variables
- Check your keys for uniqueness before joining
left_join()is your workhorse — keeps all rows from your main table- Filtering joins help identify completers vs. dropouts
- Watch for common problems: duplicate keys, different key names, NAs
- Always inspect the result to ensure it matches expectations
Before next class
📖 Read:
- R4DS Ch 18: Missing values
✅ Do:
- Submit Assignment 6
- Check if your final project needs joins
- Practice joining your own data files
The one thing to remember
If your data lives in separate files, joins are how you ask a question that spans all of them.
See you Wednesday for handling missing data!
PSY 410 | Session 13