| participant | depression_pre | depression_post | anxiety_pre | anxiety_post |
|---|---|---|---|---|
| P1 | 18 | 12 | 24 | 20 |
| P2 | 22 | 14 | 19 | 15 |
| P3 | 15 | 11 | 28 | 22 |
Data Tidying
PSY 410: Data Science for Psychology
2026-04-13
What is tidy data?
Can you plot this?
Not easily. The columns mix variables (depression vs anxiety) with time points (pre vs post). ggplot doesn’t know what to put on each axis.
By the end of today, you’ll be able to reshape this into a form that works — and understand why it needs reshaping.
The tidyverse philosophy
“Tidy datasets are all alike, but every messy dataset is messy in its own way.” — Hadley Wickham
The tools we’ve learned (ggplot2, dplyr) expect data in a specific format: tidy data.
The three rules of tidy data
- Each variable is a column
- Each observation is a row
- Each value is a cell
Simple in theory, surprisingly complex in practice.
Tidy data visualized
| participant | pre_test | post_test |
|---|---|---|
| P1 | 45 | 62 |
| P2 | 52 | 58 |
| P3 | 48 | 71 |
No! Time (pre/post) is a variable, but it’s spread across columns.
The tidy version
| participant | time | score |
|---|---|---|
| P1 | pre | 45 |
| P1 | post | 62 |
| P2 | pre | 52 |
| P2 | post | 58 |
| P3 | pre | 48 |
| P3 | post | 71 |
Now: participant, time, and score are all columns.
Why does it matter?
Tidy data works with tidyverse tools:
Wide vs. Long
Wide format
- Variables spread across columns
- One row per subject
- Humans like to read this
Long format
- Variables in a single column
- Multiple rows per subject
- R likes to work with this
Most real data needs reshaping.
Common untidy patterns
Pattern 1: Column headers are values
wide_scores <- tibble(
student = c("Alice", "Bob", "Carol"),
fall_2024 = c(85, 78, 92),
spring_2025 = c(88, 82, 95),
fall_2025 = c(91, 85, 94)
)
wide_scores# A tibble: 3 × 4
student fall_2024 spring_2025 fall_2025
<chr> <dbl> <dbl> <dbl>
1 Alice 85 88 91
2 Bob 78 82 85
3 Carol 92 95 94The semester names are values, not variable names.
Pattern 2: Multiple variables in one column
# A tibble: 3 × 2
id age_sex
<int> <chr>
1 1 25_M
2 2 32_F
3 3 28_F Age and sex are crammed into one column.
Pattern 3: Variables in rows and columns
weather <- tibble(
id = c("MX001", "MX001", "MX002", "MX002"),
year = c(2020, 2020, 2020, 2020),
month = c(1, 2, 1, 2),
element = c("tmax", "tmax", "tmin", "tmin"),
value = c(85, 87, 32, 35)
)
weather# A tibble: 4 × 5
id year month element value
<chr> <dbl> <dbl> <chr> <dbl>
1 MX001 2020 1 tmax 85
2 MX001 2020 2 tmax 87
3 MX002 2020 1 tmin 32
4 MX002 2020 2 tmin 35element contains variable names (tmax, tmin).
Psychology-specific patterns
Surveys often look like:
survey_wide <- tibble(
participant = 1:3,
bdi_1 = c(2, 1, 3),
bdi_2 = c(1, 0, 2),
bdi_3 = c(3, 2, 2),
bdi_4 = c(2, 1, 1)
)
survey_wide# A tibble: 3 × 5
participant bdi_1 bdi_2 bdi_3 bdi_4
<int> <dbl> <dbl> <dbl> <dbl>
1 1 2 1 3 2
2 2 1 0 2 1
3 3 3 2 2 1Each item is a column — wide format.
pivot_longer()
The most common tidying operation
pivot_longer() takes wide data and makes it long:
wide_scores |>
pivot_longer(
cols = fall_2024:fall_2025, # Which columns to pivot
names_to = "semester", # New column for old column names
values_to = "score" # New column for values
)# A tibble: 9 × 3
student semester score
<chr> <chr> <dbl>
1 Alice fall_2024 85
2 Alice spring_2025 88
3 Alice fall_2025 91
4 Bob fall_2024 78
5 Bob spring_2025 82
6 Bob fall_2025 85
7 Carol fall_2024 92
8 Carol spring_2025 95
9 Carol fall_2025 94Breaking it down
Selecting columns to pivot
Use any of the select() helpers:
Psychology example: Survey items
# A tibble: 12 × 3
participant item response
<int> <chr> <dbl>
1 1 bdi_1 2
2 1 bdi_2 1
3 1 bdi_3 3
4 1 bdi_4 2
5 2 bdi_1 1
6 2 bdi_2 0
7 2 bdi_3 2
8 2 bdi_4 1
9 3 bdi_1 3
10 3 bdi_2 2
11 3 bdi_3 2
12 3 bdi_4 1Now each response is its own row!
Extracting information from names
What if column names contain useful info? We want to extract the time point (t1, t2, t3).
names_prefix argument
experiment_wide |>
pivot_longer(
cols = starts_with("score"),
names_to = "time",
names_prefix = "score_", # Remove this prefix from names
values_to = "score"
)# A tibble: 9 × 3
id time score
<int> <chr> <dbl>
1 1 t1 100
2 1 t2 105
3 1 t3 108
4 2 t1 95
5 2 t2 100
6 2 t3 102
7 3 t1 110
8 3 t2 115
9 3 t3 120names_pattern argument
For more complex parsing:
names_pattern argument
For more complex parsing:
multi_scale |>
pivot_longer(
cols = -id,
names_to = c("scale", "item"),
names_pattern = "(.+)_(.+)", # Regex: anything_anything
values_to = "response"
)# A tibble: 8 × 4
id scale item response
<int> <chr> <chr> <dbl>
1 1 bdi 1 2
2 1 bdi 2 1
3 1 anxiety 1 3
4 1 anxiety 2 2
5 2 bdi 1 1
6 2 bdi 2 2
7 2 anxiety 1 2
8 2 anxiety 2 3Pivoting to calculate scale scores
Pair coding break
Your turn: 10 minutes
We defined survey_wide earlier in this deck:
# A tibble: 3 × 5
participant bdi_1 bdi_2 bdi_3 bdi_4
<int> <dbl> <dbl> <dbl> <dbl>
1 1 2 1 3 2
2 2 1 0 2 1
3 3 3 2 2 1With a partner, figure out: Which participant has the highest mean BDI score?
Tip
You’ll need pivot_longer() followed by group_by() + summarize(). The column names start with "bdi" — that’s a hint.
Before we move on
📤 Upload your code to Canvas for participation credit. Paste what you have into today’s in-class submission — it doesn’t need to work perfectly.
pivot_wider()
The opposite operation
Sometimes you need to go from long to wide:
pivot_wider() syntax
When to use pivot_wider()
- Creating summary tables for reports
- Some analyses need wide format
- Merging data that was collected differently
- Human-readable output
Multiple value columns
# Long data with multiple measures
long_multi <- tibble(
id = rep(1:2, each = 2),
time = rep(c("pre", "post"), 2),
score = c(45, 62, 52, 58),
rt = c(500, 480, 520, 490)
)
long_multi# A tibble: 4 × 4
id time score rt
<int> <chr> <dbl> <dbl>
1 1 pre 45 500
2 1 post 62 480
3 2 pre 52 520
4 2 post 58 490Multiple value columns
Separating and uniting
separate_wider_delim()
Split one column into multiple:
separate_wider_regex()
For complex patterns:
unite()
The opposite — combine columns:
Real-world examples
Example 1: Repeated measures experiment
# Data as you might receive it from SPSS
wide_rm <- tibble(
subject = 1:4,
cond_a_time1 = c(450, 520, 480, 510),
cond_a_time2 = c(420, 490, 460, 480),
cond_b_time1 = c(480, 540, 500, 530),
cond_b_time2 = c(440, 510, 470, 500)
)
wide_rm# A tibble: 4 × 5
subject cond_a_time1 cond_a_time2 cond_b_time1 cond_b_time2
<int> <dbl> <dbl> <dbl> <dbl>
1 1 450 420 480 440
2 2 520 490 540 510
3 3 480 460 500 470
4 4 510 480 530 500Tidying repeated measures
tidy_rm <- wide_rm |>
pivot_longer(
cols = -subject,
names_to = c("condition", "time"),
names_pattern = "cond_(.+)_time(.+)",
values_to = "rt"
)
tidy_rm# A tibble: 16 × 4
subject condition time rt
<int> <chr> <chr> <dbl>
1 1 a 1 450
2 1 a 2 420
3 1 b 1 480
4 1 b 2 440
5 2 a 1 520
6 2 a 2 490
7 2 b 1 540
8 2 b 2 510
9 3 a 1 480
10 3 a 2 460
11 3 b 1 500
12 3 b 2 470
13 4 a 1 510
14 4 a 2 480
15 4 b 1 530
16 4 b 2 500Now we can analyze it!
tidy_rm |>
ggplot(aes(x = time, y = rt, color = condition, group = condition)) +
stat_summary(fun = mean, geom = "point", size = 3) +
stat_summary(fun = mean, geom = "line") +
stat_summary(fun.data = mean_se, geom = "errorbar", width = 0.1) +
labs(
title = "Reaction Time by Condition and Time",
x = "Time Point",
y = "Reaction Time (ms)"
) +
theme_minimal(base_size = 14)Now we can analyze it!
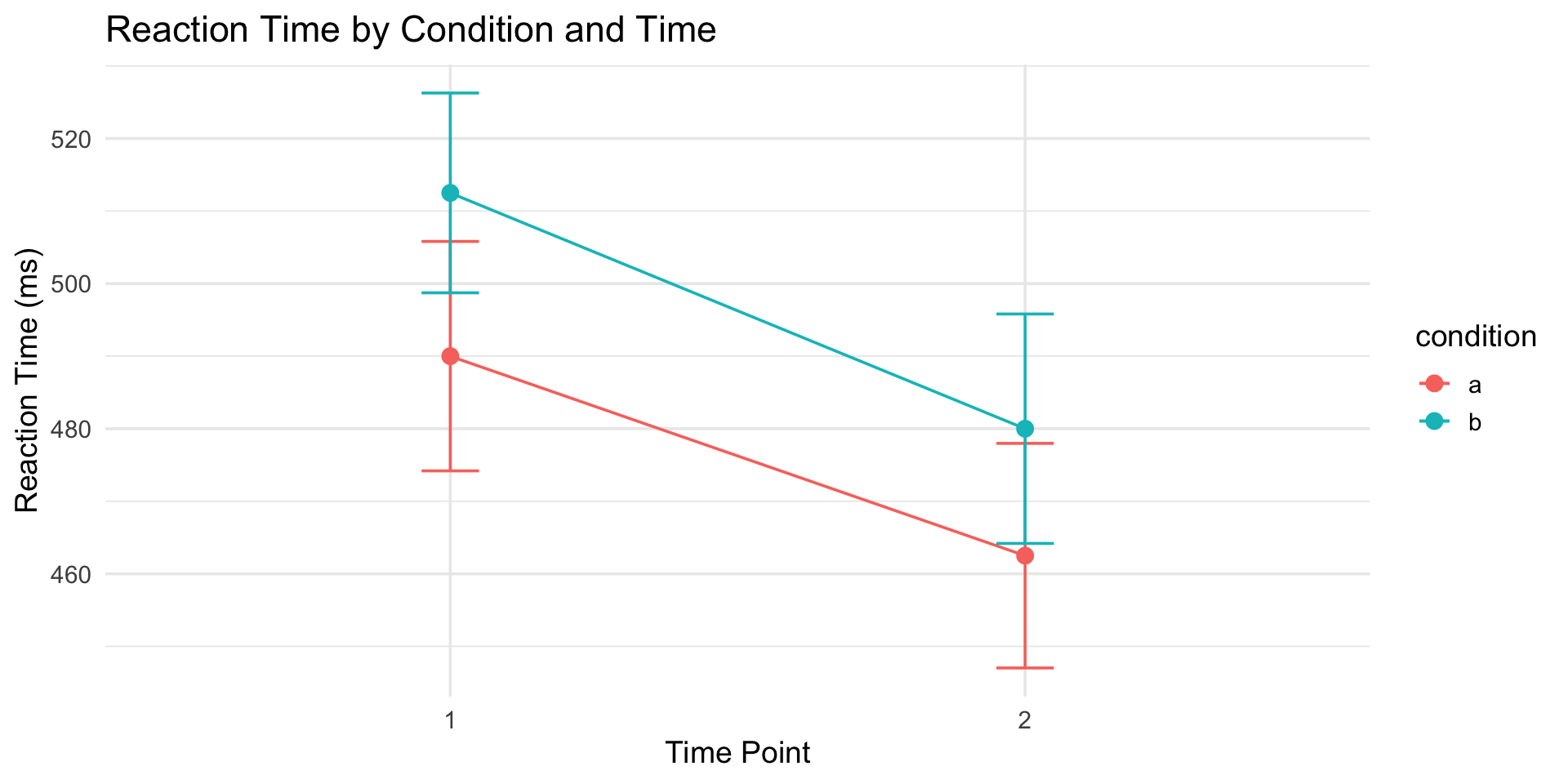
Example 2: Questionnaire with subscales
# Raw questionnaire data
quest <- tibble(
pid = 1:3,
anx_1 = c(3, 2, 4), anx_2 = c(2, 3, 3), anx_3 = c(4, 2, 5),
dep_1 = c(2, 1, 3), dep_2 = c(3, 2, 4), dep_3 = c(2, 1, 3)
)
quest# A tibble: 3 × 7
pid anx_1 anx_2 anx_3 dep_1 dep_2 dep_3
<int> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 1 3 2 4 2 3 2
2 2 2 3 2 1 2 1
3 3 4 3 5 3 4 3Example 2: Questionnaire with subscales
# Tidy and calculate subscales
quest |>
pivot_longer(
cols = -pid,
names_to = c("scale", "item"),
names_pattern = "(.+)_(.+)",
values_to = "response"
) |>
group_by(pid, scale) |>
summarize(subscale_mean = mean(response), .groups = "drop") |>
pivot_wider(names_from = scale, values_from = subscale_mean)Example 2: Questionnaire with subscales
# A tibble: 3 × 3
pid anx dep
<int> <dbl> <dbl>
1 1 3 2.33
2 2 2.33 1.33
3 3 4 3.33Example 3: Multilevel/nested data
# Students nested in classrooms
students <- tibble(
classroom = rep(c("A", "B"), each = 3),
student = 1:6,
pretest = c(70, 75, 72, 68, 71, 69),
posttest = c(80, 82, 78, 75, 79, 77)
)
students# A tibble: 6 × 4
classroom student pretest posttest
<chr> <int> <dbl> <dbl>
1 A 1 70 80
2 A 2 75 82
3 A 3 72 78
4 B 4 68 75
5 B 5 71 79
6 B 6 69 77Example 3: Multilevel/nested data
# Tidy for analysis
students |>
pivot_longer(
cols = c(pretest, posttest),
names_to = "time",
values_to = "score"
)# A tibble: 12 × 4
classroom student time score
<chr> <int> <chr> <dbl>
1 A 1 pretest 70
2 A 1 posttest 80
3 A 2 pretest 75
4 A 2 posttest 82
5 A 3 pretest 72
6 A 3 posttest 78
7 B 4 pretest 68
8 B 4 posttest 75
9 B 5 pretest 71
10 B 5 posttest 79
11 B 6 pretest 69
12 B 6 posttest 77pivot_longer() trips everyone up the first time
Pitfall 1: Forgetting what’s a variable
Ask yourself: What are my variables?
- Participant ID? ✓ Variable
- Time point? ✓ Variable (not separate columns!)
- Score? ✓ Variable
- Item number? Depends on your analysis
Pitfall 2: Over-pivoting
Not everything needs to be long:
Age, gender, and score are different variables — keep them as columns.
General tidying strategy
- Identify the variables (what are you measuring?)
- Look at your current structure (what’s a row? column?)
- Determine what operations you need
- Test with a small subset first
- Verify you haven’t lost data
Get a head start
Try it yourself
We created wide_rm earlier — a repeated measures dataset:
On your own:
- Pivot it to long format, extracting condition and time from the column names
- Calculate mean RT by condition and time
- Sketch (on paper or in ggplot) what you’d expect the plot to look like
This is very close to what Assignment 3 will ask you to do.
Wrapping up
The tidyr toolkit
| Function | What it does |
|---|---|
pivot_longer() |
Wide → Long |
pivot_wider() |
Long → Wide |
separate_*() |
Split columns |
unite() |
Combine columns |
The tidy data mantra
- Each variable is a column
- Each observation is a row
- Each value is a cell
When in doubt, ask: “What would make this easiest to plot/analyze?”
Before next class
📖 Read:
✅ Practice:
- Reshape a dataset you’ve worked with
- Try tidying some messy example data
- Practice
pivot_longer()— it’s the most common
Key takeaways
- Tidy data has a specific structure that works with tidyverse
- pivot_longer() is your most-used tidying function
- pivot_wider() is useful for tables and some analyses
- Think about your variables before reshaping
- Column names contain information — extract it with
names_pattern
The one thing to remember
If your data isn’t tidy, your analysis can’t start. pivot_longer() is the bridge.
Next time: Data Import
PSY 410 | Session 5